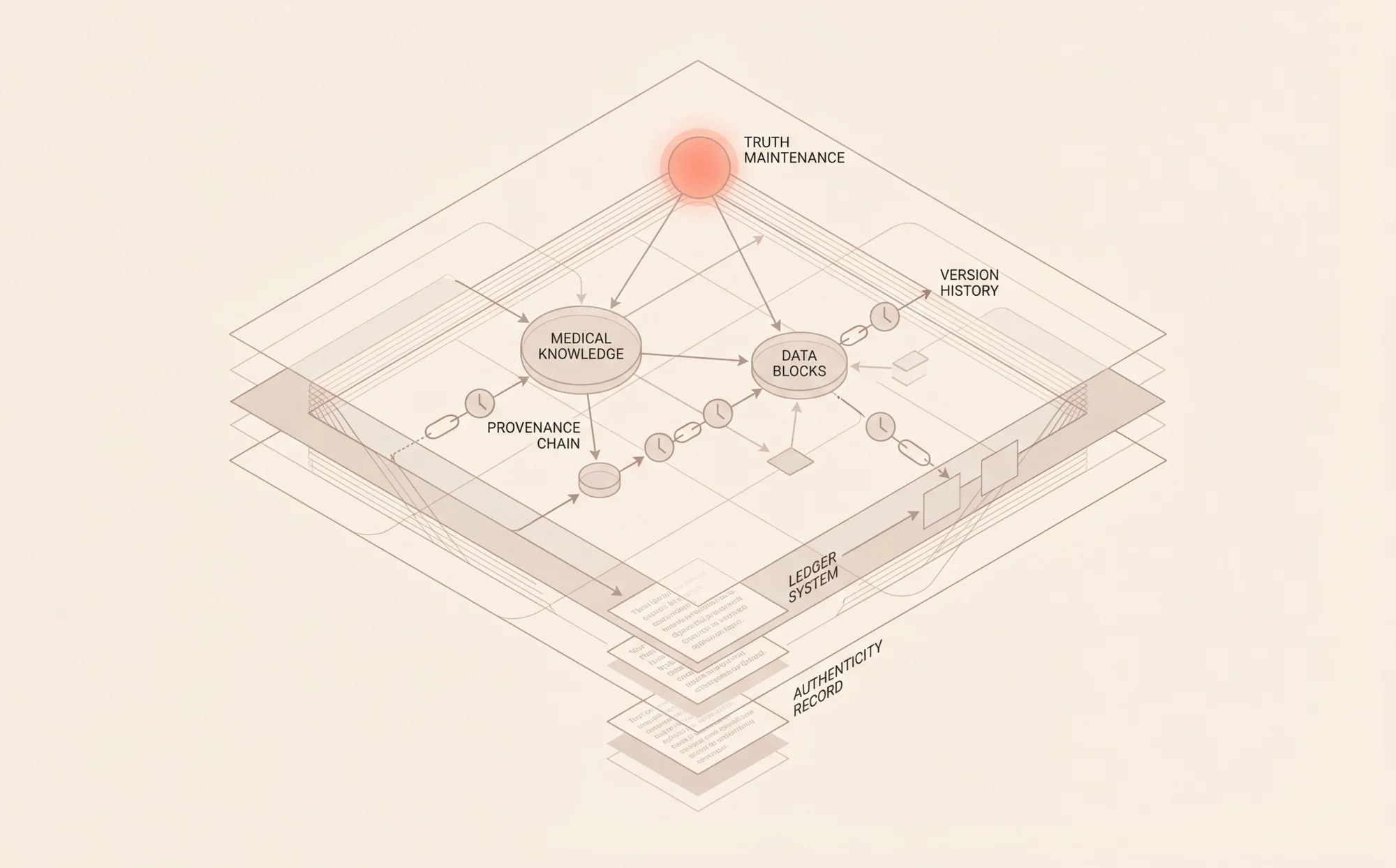
Building a Medical Memory Ledger: Provenance, Truth Maintenance, and Revocation
1. The Imperative for Structured Memory Systems in Healthcare
The prevailing paradigm for AI agent memory—an ever-expanding, unstructured vector store or conversation buffer—is fundamentally inadequate for high-stakes domains like healthcare. These conventional memory systems in healthcare lack the structure, provenance, and mechanisms for correction or revocation essential for clinical safety. In a clinical context, where patient safety and healthcare data integrity are paramount, an agent must not only recall a fact but also understand its origin—the core of provenance in clinical data—its temporal validity, and the protocol for its amendment or retraction. A clinical agent operating with an unmanaged memory model presents an unacceptable risk of propagating erroneous or outdated information, potentially leading to adverse patient outcomes.
This technical deep dive addresses this critical gap by detailing the architecture and implementation of a Medical Memory Ledger. This system treats agent memory not as a simple log but as a structured, auditable database with integrated truth maintenance. We will explore a relational-graph hybrid model where every stored fact is a node with typed provenance edges, linking it to source documents, timestamps, confidence scores, and originating agents. The scope of this guide covers schema design, the implementation of a Justification-Based Truth Maintenance System (JTMS), conflict resolution engines, consent-aware revocation protocols, and the generation of audit-ready provenance chains to ensure clinical data accountability.
Technical Prerequisites:
- Proficiency in Python (version 3.10+).
- Experience with graph databases (e.g., Neo4j, ArangoDB) and their query languages (e.g., Cypher, AQL).
- Familiarity with data modeling using Pydantic or similar libraries.
- Conceptual understanding of knowledge representation and reasoning systems.
Upon completing this guide, you will be equipped to design and implement a robust, provenance-tracked memory system capable of managing the lifecycle of clinical data with the rigor required for healthcare applications.
2. Core Architecture: A Hybrid Model for Healthcare Data Integrity
The foundation of this patient data provenance system is a hybrid data architecture that combines the rich, queryable attributes of a relational model with the expressive, interconnected nature of a graph database. This design allows us to store facts with detailed metadata while explicitly modeling the complex relationships of derivation and justification between them.
The Fact Node: The Atomic Unit of Memory
Each piece of information, whether a diagnosis, a vital sign, or an allergy, is encapsulated as a Fact Node. This node is more than a simple statement; it is a structured object containing the core data payload alongside critical metadata. These attributes include a unique identifier, the fact's content (e.g., "Patient has an allergy to penicillin"), a confidence score, a status (e.g., CURRENT, REVOKED, CORRECTED), and timestamps for creation and modification.
Provenance Edges: Linking Facts to Origins
The graph nature of the ledger is realized through Provenance Edges. These are directed, typed edges that connect Fact Nodes to their sources or to other Fact Nodes from which they were derived. An edge might be of type EXTRACTED_FROM, linking a Fact Node to a SourceDocument node, or of type INFERRED_FROM, linking a new diagnosis to the lab results that support it. Each edge also carries metadata, such as the ID of the agent or process that created the link and the exact timestamp of its creation.
Component Interactions
The system comprises three primary interacting components. The Ingestion Pipeline processes incoming data, creates Fact Nodes and Provenance Edges, and persists them. The Truth Maintenance System (TMS) monitors the graph for changes—such as the retraction of a source document—and propagates belief updates throughout the network of dependent facts. Finally, the Query Engine provides an API for the clinical agent to retrieve information, resolve conflicting data, and generate complete audit trails for any given fact.
3. Designing Patient-Centric Data Models for the Ledger Schema
A precise and extensible schema is crucial for achieving high healthcare data integrity. We recommend using a library like Pydantic to define and enforce the structure of our nodes and edges, ensuring data consistency from the point of ingestion.
Defining the Fact Node Schema
The FactNode serves as the core entity. Its schema must capture not only the clinical data but also the metadata required for truth maintenance and auditing.
This schema provides a robust structure. The content field is a flexible dictionary to accommodate various patient-centric data models, ideally aligned with standards like HL7 FHIR. The justifications list is key for the TMS, forming the explicit dependency graph.
Modeling Provenance Edges
While justifications are stored on the node, the explicit graph edges model the direct lineage. The edge schema captures the nature of the relationship.
Integrating Clinical Ontologies
For the content field to be computationally useful, it must be normalized against standard clinical ontologies. We recommend using coding systems like SNOMED CT for diagnoses and procedures, LOINC for lab results, and RxNorm for medications. This normalization enables semantic queries and more intelligent conflict resolution.
4. Implementation: Ingestion and Tracking Provenance in Clinical Data
The ingestion pipeline is the entry point for all data into the Medical Memory Ledger. It is responsible for parsing, structuring, and linking new information correctly.
Here is a step-by-step guide to the ingestion process:
- Receive Data: An API endpoint receives a data payload, which could be a structured FHIR resource, an unstructured clinical note, or a patient-submitted correction.
- Generate Source Node: A unique
SourceDocumentnode is created in the graph. A hash of the document content is computed and stored to prevent duplicate processing and to anchor the chain of provenance in clinical data. - Extract Facts: An NLP model or a structured data parser iterates through the document to identify discrete pieces of clinical information. A 2021 study by the Journal of Medical Internet Research found that up to 15% of patient data in unstructured clinical notes contains contradictions, highlighting the need for discrete fact extraction.
- Create Fact Nodes: For each extracted piece of information, a
FactNodeinstance is created according to the schema defined previously. Thesource_hashandagent_idare populated. - Establish Provenance: A
ProvenanceEdgeof typeEXTRACTED_FROMis created, linking each newFactNodeto theSourceDocumentnode. - Persist to Database: All new nodes and edges are committed to the graph database in a single transaction to ensure atomicity.
The following code demonstrates a simplified ingestion function.
An API call to trigger this process might look like this:
5. Truth Maintenance: Ensuring Logical Consistency and Data Integrity
A Truth Maintenance System is the reasoning engine that maintains the logical consistency that underpins healthcare data integrity. When a base fact is altered or retracted, the TMS is responsible for propagating this change to all dependent facts, a key aspect of our revocation rules for health data. We will focus on a Justification-Based TMS (JTMS), which is well-suited for this architecture.
Core JTMS Concepts
In a JTMS, each fact (or node) has a belief status, typically IN (believed to be true) or OUT (not believed to be true). The status of a fact is determined by its justifications. A justification is a record that states a fact is valid if certain other facts (its antecedents) are IN. When an antecedent's status changes, the TMS re-evaluates the status of all facts that depend on it.
A Simplified JTMS Engine
The following Python code illustrates a basic JTMS engine that can be integrated with the graph database. It manages belief states and propagates retractions.
In a production system, a change to a FactNode's status in the graph database would trigger a function that initiates this belief propagation process, updating the status field of affected nodes throughout the graph.
6. The Conflict Resolution Engine: A Key to Data Accuracy
Clinical data often contains contradictions. A patient's record might list an allergy in one note but not in another, or two different clinicians might provide conflicting diagnoses. The Medical Memory Ledger requires a deterministic conflict resolution engine to present the most reliable information to the AI agent and maintain healthcare data integrity.
Our engine resolves conflicts using a weighted scoring model based on three primary factors:
- Source Authority: Different sources have different levels of reliability. A diagnosis from a board-certified specialist is weighted more heavily than a self-reported symptom from a patient intake form.
- Recency: More recent information is generally considered more accurate, especially for time-sensitive data like vital signs or active prescriptions.
- Domain Hierarchy: In some cases, a more specific fact supersedes a general one (e.g., "acute bronchitis" is more specific than "respiratory infection").
The following code provides a function to resolve a conflict between two FactNode objects.
When the query engine detects multiple facts concerning the same clinical concept (e.g., two different allergy statuses for penicillin), it invokes this resolver to determine which fact should be considered CURRENT. The losing fact's status is then updated to OUTDATED.
7. Data Lifecycle: TTL Policies in Clinical Records and Consent-Aware Revocation
The lifecycle of clinical data is complex. Some facts have a short period of relevance, while others must be purged based on patient consent.
Implementing TTL Policies
Not all medical data remains relevant indefinitely. A patient's heart rate from three years ago is likely irrelevant, whereas a documented allergy is permanent. The ledger supports the full data lifecycle with TTL policies in clinical records by setting the ttl_seconds field on a FactNode.
A background process or a database-native TTL feature can then be used to manage these facts.
Consent-Aware Revocation
Patient consent is a cornerstone of healthcare data management. If a patient withdraws consent for a specific document or data source to be used, the system must not only delete the source but also purge all facts derived from it, enforcing strict revocation rules for health data. This is where the provenance graph is indispensable.
The revocation process involves a graph traversal, starting from the source node whose consent has been withdrawn.
This ensures that a consent withdrawal is fully honored, purging all derived and inferred knowledge from the agent's memory.
8. Enhancing Trust: Patient Corrections and Clinical Data Accountability
Achieving clinical data accountability requires transparency and mechanisms for correction. The ledger is designed to support both.
Designing a Patient Correction Workflow
When a patient identifies an error in their record, they can flag a specific FactNode. This action does not immediately delete the fact but instead initiates a workflow:
- The
FactNode's status is changed toNEEDS_REVIEW. - A new
FactNodeis created with the patient's corrected information, linked to the original with aCORRECTSedge. - A notification is sent to a human clinician for review.
- The clinician reviews the evidence and either accepts the correction (making the new fact
CURRENTand the old oneCORRECTED) or rejects it (reverting the original fact toCURRENT).
Generating Audit-Ready Trace Logs
The ledger's explicit provenance graph makes auditing straightforward. For any decision made by the AI agent, we can reconstruct the complete provenance chain, ensuring full clinical data accountability.
This functionality is critical for explainability (XAI) and for regulatory compliance, as it provides a verifiable record of the agent's reasoning process.
9. Best Practices for Performance and Scalability
Deploying a Medical Memory Ledger at scale requires careful attention to performance and reliability. We recommend the following best practices:
- Database Indexing: Create indexes on frequently queried attributes, such as
fact_id,source_hash, and fields within thecontentdictionary (e.g., SNOMED codes). For graph traversals, ensure the database is optimized for the specific patterns used by the TMS and audit queries. - Caching Strategies: Implement caching strategies (e.g., Redis) for frequently accessed, stable facts. This can significantly reduce read latency for the AI agent, but cache invalidation must be tightly coupled with the TMS to avoid serving stale data.
- Asynchronous Processing: TMS belief propagation can be computationally intensive. Offload these updates to a background worker queue (e.g., Celery, RabbitMQ) to keep the ingestion API responsive. The agent can be notified of belief changes via websockets or a similar mechanism.
- Optimize Graph Traversal: Write graph queries to be as specific as possible, limiting the depth and breadth of traversals. Use edge direction and type to prune the search space during provenance tracing and revocation cascades.
10. Troubleshooting Common Issues in Medical Memory Systems
Implementing a system with this level of complexity can present unique challenges. Below are common issues and their solutions.
-
Problem: Circular Dependencies in TMS
- Symptom: A belief propagation update enters an infinite loop, causing high CPU usage. This occurs if Fact A justifies Fact B, and Fact B justifies Fact A.
- Solution: Implement cycle detection in your
_update_beliefpropagation logic. Before traversing to a dependent, check if it is already in the current update path's call stack. If so, break the loop and flag the cycle for manual review.
-
Problem: Belief "Flapping"
- Symptom: A fact's status rapidly oscillates between
INandOUTdue to high-frequency, conflicting updates from different sources. - Solution: Introduce a debouncing mechanism. When a fact's belief status changes, wait for a short stabilization period (e.g., 500ms) before propagating the change. Aggregate all changes within this window and process them as a single update.
- Symptom: A fact's status rapidly oscillates between
-
Problem: Inconsistent State After a Crash
- Symptom: A system failure during a TMS update leaves the graph in a logically inconsistent state.
- Solution: Ensure all TMS propagation runs are transactional. The entire cascade of status updates should be committed to the database as a single atomic operation. If the transaction fails, the database should roll back to the last known consistent state.
11. API Reference
A well-defined API is essential for integrating the Medical Memory Ledger with other clinical systems and AI agents.
Key API Endpoints
| Method | Endpoint | Description |
|---|---|---|
| POST | /v1/ingest | Ingests a new source document and extracts facts. |
| GET | /v1/query/fact/{fact_id} | Retrieves a specific fact node by its ID. |
| POST | /v1/query/semantic | Queries for facts based on clinical codes and temporal filters. |
| POST | /v1/revoke | Initiates a consent-aware revocation cascade for a given source ID. |
| POST | /v1/correct | Submits a patient- or clinician-initiated correction for a fact. |
| GET | /v1/audit/trace/{fact_id} | Retrieves the full provenance chain for a given fact. |
12. Conclusion: A New Standard for Clinical Data Accountability
The Medical Memory Ledger represents a paradigm shift from simplistic, unstructured agent memory to a robust, auditable, and logically consistent knowledge base. By combining patient-centric data models in a relational-graph structure with a truth maintenance system, we create a framework that respects the complexities of clinical data—its provenance, its temporal nature, and the critical importance of correction and consent. This architecture is not merely a technical improvement; it is a foundational requirement for deploying AI agents safely and responsibly in the healthcare domain.
Designing these advanced patient data provenance systems is key to the future of AI in medicine. As regulations evolve, the ability to enforce revocation rules for health data and demonstrate healthcare data integrity will become non-negotiable. This ledger provides the blueprint for that future.
For further exploration, your team should consult research on Assumption-Based Truth Maintenance Systems (ATMS) for handling multiple contexts, as well as production-grade graph database documentation for advanced optimization techniques. The next steps for a production deployment involve rigorous testing of the TMS under concurrent load, integration with clinical authentication and authorization systems, and establishing formal data governance policies for the ledger.